TECHNOLOGY
WHAT IS DNA-PAINT?
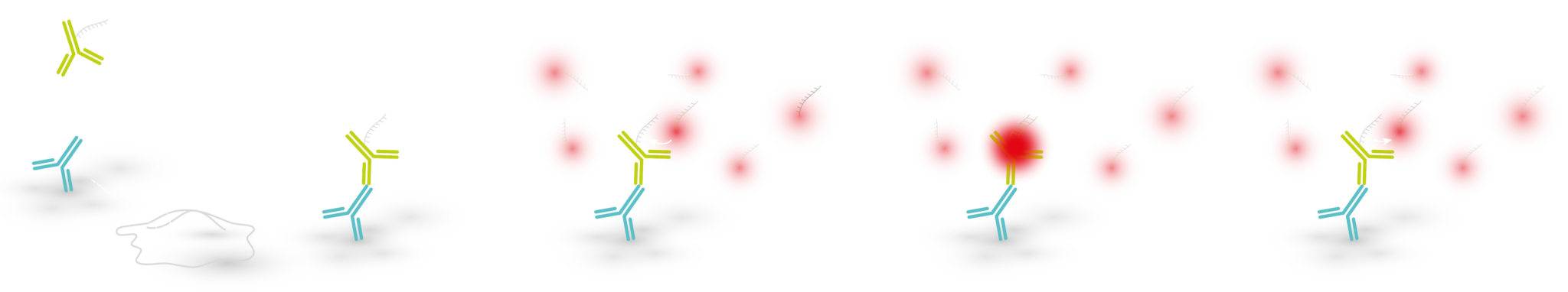
DNA-PAINT (DNA-Point Accumulation for Imaging in Nanoscale Topography) is a localization based super-resolution microscopy method that uses short, dye-labeled single stranded DNA (ssDNA strands) that transiently bind to their complements:
- Imager strand – a dye-labeled DNA oligo present in imaging buffer
- Docking strand – a DNA oligo complementary to the imager strand that is delivered to the target of interest via binders (e.g. antibodies, single domain antibodies)
WHY USE DNA-PAINT?
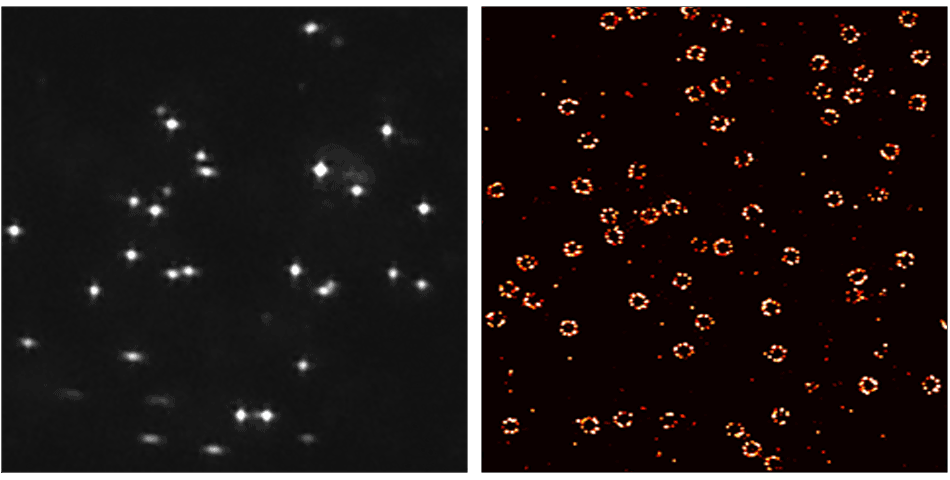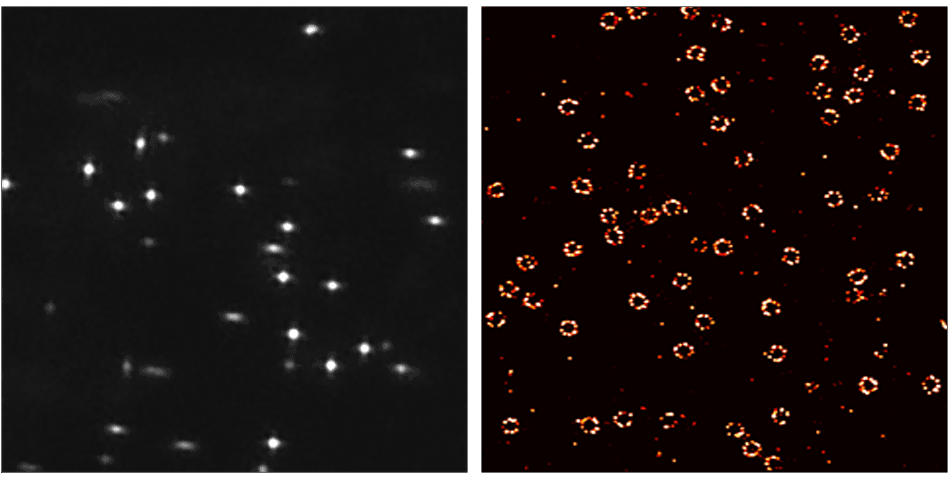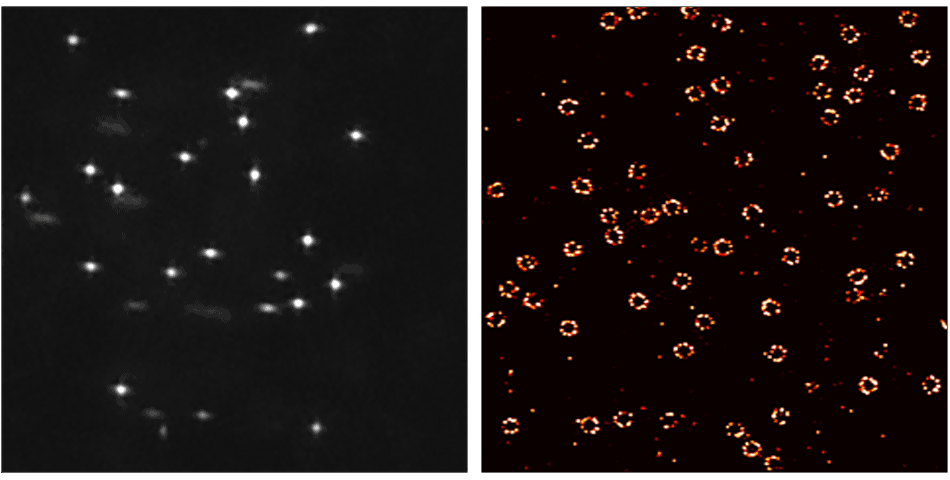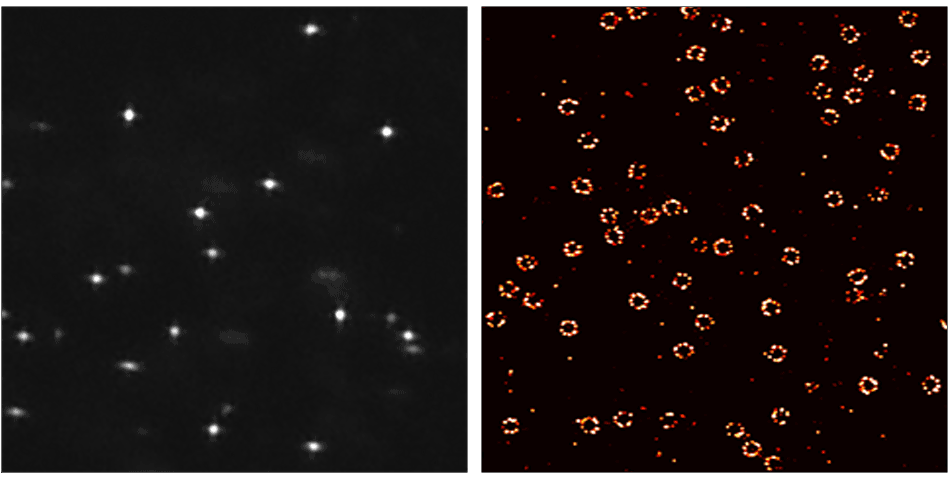
- Ultra resolution! Resolve 10 – 15 nm distances in cells.
- Unlimited target multiplexing! In DNA-PAINT, spectrally compatible dyes aren’t needed to image multiple targets at a time. Instead, you can simply couple different docking strands (e. different DNA sequences) to your target binders and image each target sequentially using the same laser line with its corresponding imager strand.
- Target Counting! The predictable nature of DNA binding kinetics allows you to quantify how many proteins are present in cells (qPAINT).
WHERE TO START?
- Simple TIRF Microscope – like the ZEISS Elyra 7, Nikon N-STORM, ONI Nanoimager, abbelight SAFe 180, Bruker VUTARA VXL microscope, Andor Dragonfly or similar system found in your imaging facility.
- DNA-PAINT compatible binder – any antibody or single-domain antibody that binds to your target protein and is coupled to a docking strand. Find a DNA-PAINT compatible binder here at Massive Photonics! We have a wide range of antibodies and single domain antibodies. Order your custom antibody or single-domain antibody (PRODUCTS page).
- Picasso data analysis software – a free software developed for DNA-PAINT image analysis (Download Picasso at Github). Find out how to use Picasso in Nature Protocols 12, 1198–1228 (2017)
RESI
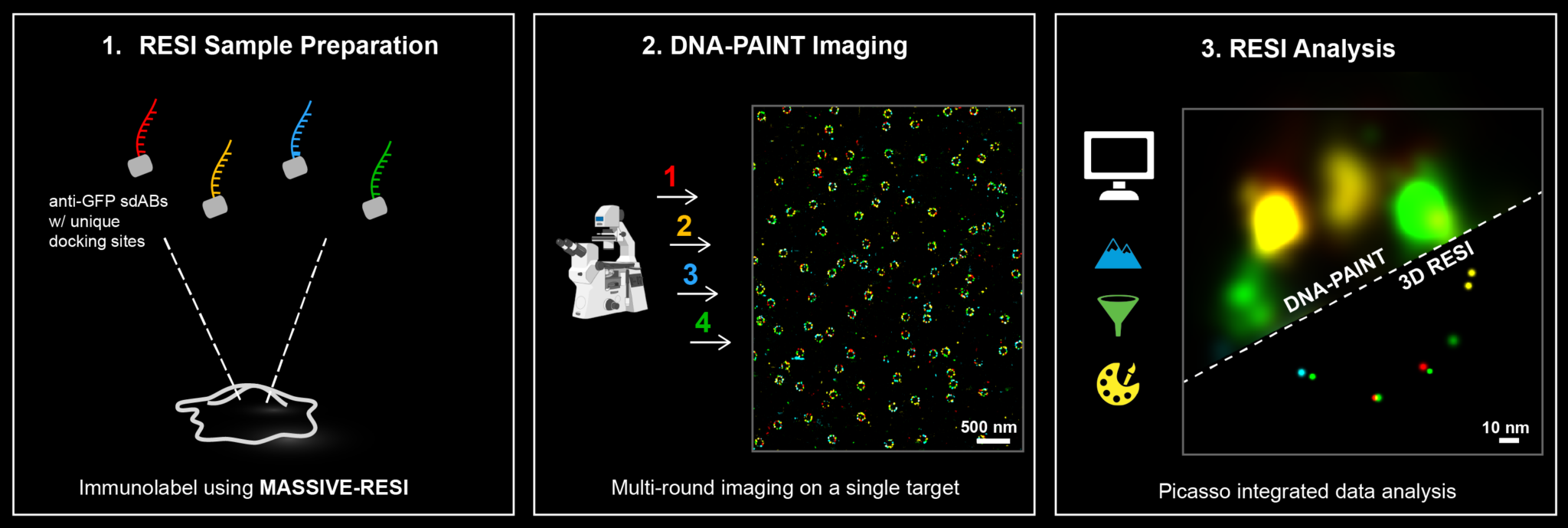
WHAT IS RESI?
Resolution Enhancement by Sequential Imaging (RESI) is a paradigm shifting DNA-PAINT super-resolution microscopy technique that enables sub nanometer resolution (Nature 617, 711–716 (2023)).
How does RESI work?
RESI takes advantage of the ability of DNA-PAINT to encode target identities through DNA sequences. To visualize multiple targets in a single DNA-PAINT experiment, called multiplexing, each target is imaged sequentially with orthogonal docking and imager strands. RESI uses the same multiplexing principle for a single target species, creating sparser and more isolated localizations. In each imaging round, the center of each localization group is determined to create a resolution enhancement. Resolution enhancement scales with the number of imaging rounds performed.
Experimentally, RESI can be easily performed by immunolabeling a target species with the same single-domain antibody binder conjugated to orthogonal DNA-PAINT docking sequences. The sample then undergoes multiple imaging and washing rounds until all sequences are imaged.